State-of-the-art integrated genomics and bioinformatics services
The Genomic Technologies Core Facility incorporates bioinformatics analyses to facilitate a complete service from experimental design through to data analyses.
We provide fast, affordable and reliable services for the most popular, cutting-edge applications of genomic technologies.
With a seamless flow to bioinformatics analyses, we can offer end-to-end support for research projects encompassing initial planning and experimental design, through to sample processing and data analyses.
In addition to The University of Manchester’s researchers, we also support external access.
How we work
Workflow and projects
We provide an end-to-end service, from samples to results.
Our workflow
Our operating model for service provision is one of ‘sample-in, data-out’. This means users simply submit their nucleic acid samples and the facilities do the rest.
The whole workflow involves various levels of quality control (QC) and processing steps and culminates with bioinformatics analyses.
Some of our platforms are also available for use directly by trained researchers.

Projects
We work on projects that range from studying basic biological mechanisms such as developmental processes and immune system function, through to understanding disease through uncovering deregulated pathways, disease signatures and heterogeneity at the single cell level.
Applications
Experience and techniques
We have extensive experience in a wide range of applications to support researchers from across the Faculty of Biology, Medicine and Health and beyond.
Typical applications include:
Transcriptomics
Transcriptomics and differential expression experiments, for example RNA-seq.
Epigenomic analyses
Epigenomic analyses such as ChiP-seq and ATAC-seq.
Single-cell studies
Single-cell studies such as scRNA- and scATAC-seq, and spatial transcriptomics.

Targeted approaches
More targeted approaches addressing specific biological questions.
Technologies and equipment
Supported technologies
We invest in an extensive range of technology platforms and equipment to provide the most up-to-date techniques.
The facility occupies purpose-built laboratory space that supports four main technology categories:
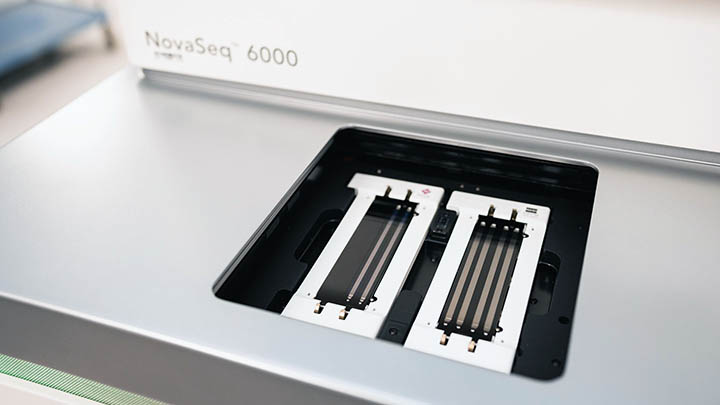
Comprehensive next-generation sequencing (NGS) solutions including library preparation, sequencing systems, and data analysis.
Illumina sequencing enables a wide variety of applications, allowing researchers to ask virtually any question related to the genome, transcriptome, or epigenome of any organism.
Find out more on the Illumina sequencing webpage.
The majority of our Illumina sequencing is performed on a NovaSeq6000, although we also have iSeq and MiniSeq platforms available for quality control and development purposes for smaller projects.
A useful tool to compare sequencing platforms can be found on the Illumina website.
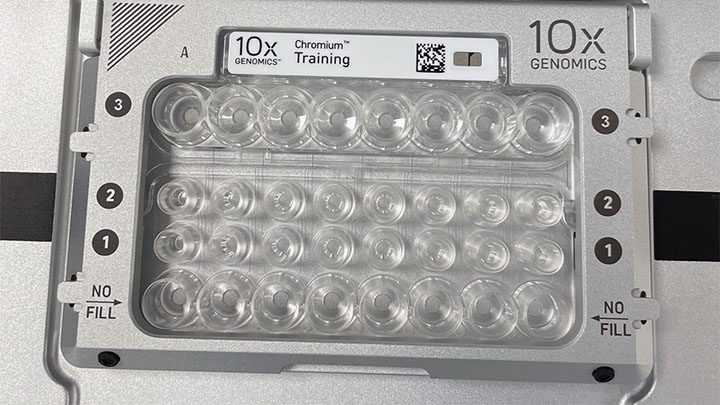
Enabling the study of cellular heterogeneity found in biological systems. Primarily using the technology and equipment of 10X Genomics, these approaches are providing insights into the cell types found in healthy and diseased tissues in humans and during development of model organisms.
Find out more on our Single Cell website.
StepOnePlus 96-well Real-Time PCR instrument
Perfect for both first-time and experienced users.
Find out more on the Thermofisher website.
QuantStudio12K Flex
The all-in-one qPCR instrument. Maximum throughput, ultimate flexibility.
Find out more on the Thermofisher website.
BioMarkHD High-definition biology
Flexible, sensitive, fast and efficient.
QX200 ddPCR
Bio-Rad’s second-generation digital PCR system provides absolute quantification of target DNA or RNA molecules using EvaGreen or TaqMan Hydrolysis Probes. Unmatched sensitivity and precision for various applications.
Find out more on the BioRad website.
The nCounter® platform provides a simple and cost-effective solution for multiplex analysis of up to 800 RNA, DNA, or protein targets. This platform accelerates research, requiring just 15 minutes total hands-on time and no amplification, cDNA conversion, or library prep, generating publication ready figures in roughly 24 hours.
Find out more on the NanoString website.
Publications and outputs
Supporting high impact publications
The Genomics Technologies core facility has been instrumental in research leading to top-class outputs.
Here is a small selection of key publications we have contributed to.
Nucleic Acids Research, 2019.
We developed a novel integrated bioinformatics pipeline for single cell ATAC-seq data analysis, and demonstrated its utility for separating cancer cell populations based on data generated in the Genomic Technologies core facility.
- DOI: 10.1093/nar/gky950
Nature Communications, 2020.
This study generated replicated ChIP-seq of three Histone marks in 13 human embryonic tissues between weeks 5 and 8 of development. We described patterns of promoter and enhancer usage across the tissues and tested tissue-specificity and functional impact of a number of human embryonic enhancers.
Proceedings of the National Academy of Sciences of the United States of America, 2020.
In this paper, integration of RNAseq and antibody-independent ChIP-seq datasets from locally generated CRISPR-edited mice revealed a new role for hepatic REVERBa. REVERBα serves to buffer against mistimed metabolic challenge rather than drive basal rhythmicity in metabolic activity.
Nature Communications, 2020.
A comprehensive range of genomic technologies including bulk RNA-seq, exome sequencing and single cell transcriptomics were used to facilitate the establishment of a living biobank of ovarian cancer ex vivo models revealing profound mitotic heterogeneity.
eLife, 2020.
This study identified a novel role for KLF5 in cell cycle control in cancer and was based on ATAC-seq, RNA-seq and ChIP-seq data which revealed how KLF5 transcription factor activity is altered in cancer cells.
- DOI: 10.7554/eLife.57189
External access
Working with industry and other academic institutions
Although primarily focused on supporting research at The University of Manchester, we are also able to support work from other academic institutions and industry.
Academic institutions
Our capacity for performing external work is dependent on the availability and current usage levels of the platforms required, and also on the complexity of the work to be performed.
Access to the GTCF services for experienced researchers who can prepare their own ‘ready-to-process’ samples can be especially straight forward. Our normal turnaround time for both these and our full-service support can be up to six weeks, depending on the required technology.
The NanoString nCounter and Fluidigm BioMarkHD are prime examples of technology platforms where we have the capability and with which we are perfectly placed to service the scientific requirements of researchers in this field
Please contact the Facility Manager for further details:
Dr Andy Hayes
Email: andy.hayes@manchester.ac.uk

Industry
We have worked for a number of multinational companies, both on an individual basis and as part of packages of work involving multiple facilities. Work may be on a collaborative basis or purely as a service provision.
All work is fully documented and contractual, and non-disclosure compliance is assured.
If you are interested in accessing the facility please contact our Business Development Manager for further details:
Dr Joanne Flannelly
Email: joanne.flannelly@manchester.ac.uk

Contact us
Find out more
Get in touch for further information or to inquire about using our facility.
Dr Andy Hayes, Senior Experimental Officer and Facility Manager
Email: andy.hayes@manchester.ac.uk
Tel (office): +44 (0)161 275 1589
Tel (lab): +44 (0)161 275 1551
Genomic Technologies Core Facility
Faculty of Biology, Medicine and Health
D.1528 Michael Smith Building
The University of Manchester
M13 9PT
Maps and travel
We are based in the AV Hill Building (Building 75 on the University campus map).
The facility is housed in a purpose-built laboratory in room G.006.
Technology platforms
Technology platforms
We have a pioneering environment and facilities for research, innovation and technology development.
Technology platforms main page
