Access, training and support for imaging systems
The Bioimaging Core Facility provides access, training and support for state-of-the-art AFM, widefield, confocal and high-content live cell imaging.
We provide access to a wide range of modern light microscope systems from Atomic Force Microscopy (AFM) through to in vivo multi-photon confocals.
This means we have a microscope for almost every imaging application and are happy to discuss your requirements and arrange training/access on the most appropriate systems.
In addition to The University of Manchester’s researchers, we also support external access.
How we work
Supporting your imaging project
We give support throughout your imaging project from experiment design through to image acquisition, processing and data analysis.
Microscopy projects
We encourage new users to discuss their microscopy projects with us before initiating work.
This allows us to advise on experiment design, sample preparation and to discuss the various imaging options.
Once we have agreed on the best system/s to use we provide full and ongoing user training. This ensures that all our users can use the systems independently and to their full potential.
Slidescanning service
We also offer a slidescanning service where the slides are automatically and fully digitised in either colour brightfield or up to four fluorescent channels.

Applications
How we can help
We have a wide range of microscope systems that offer advanced imaging capabilities.
If you have an imaging project, even if it’s new and novel, then it is likely that we have at least one microscope that can meet your requirements. If it isn’t available, we’ll work with you to make it available.
Typical experiments include:
Live cell imaging
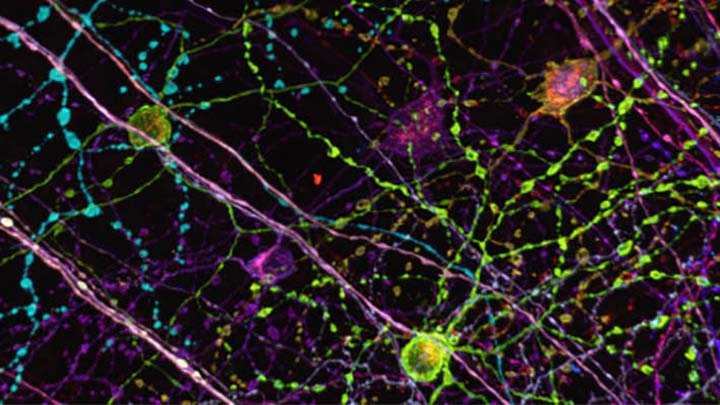
Live cell imaging of cells over 1 to 72 hours using widefield, spinning disc or point scanning confocals to allow tracking and protein expression studies.
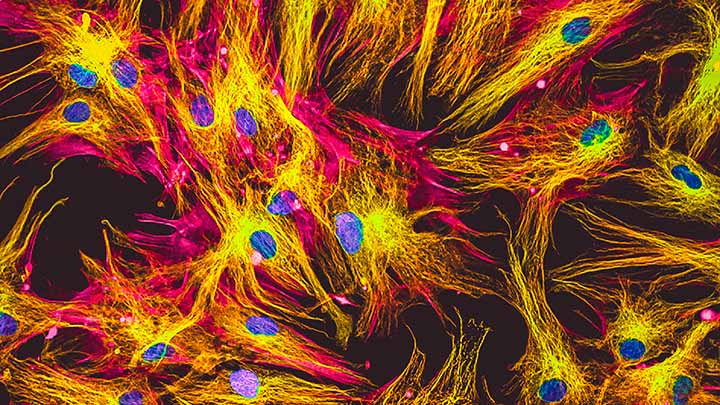
Fixed cells and tissue
Colocalisation and expression patterns of fixed cells and tissue.
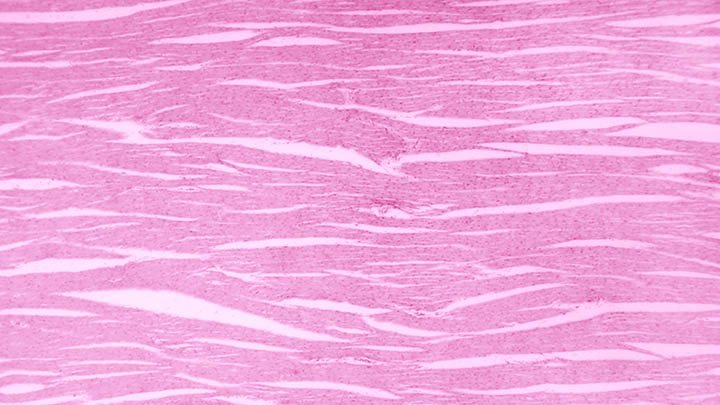
Protein-protein
Protein-protein interactions investigated using FCS, FLIM and super resolution microscopy.

Protein dynamics
Protein dynamics studied using photokinetic events such as FRAP, FLIP and photoactivation.
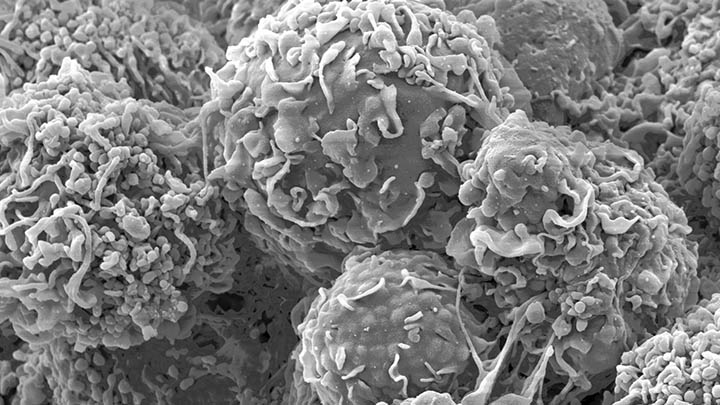
Structural characterisation and mechanical sensing
Structural characterisation and mechanical sensing using biosensors (FLIPPER and FRET/FLIM).
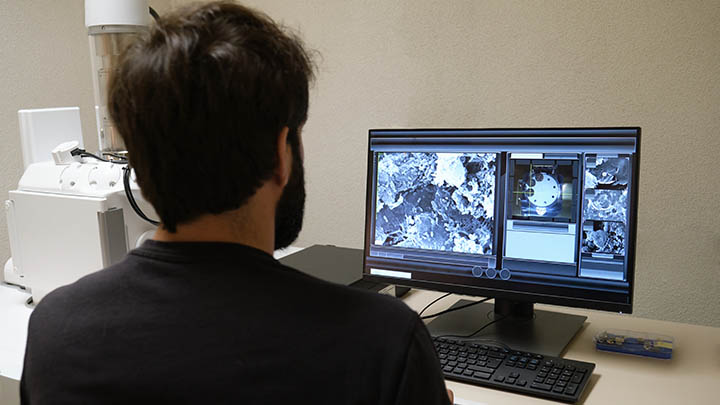

Ultra-structural properties
Ultra-structural properties of cell and tissue using Atomic Force Microscopy (AFM).
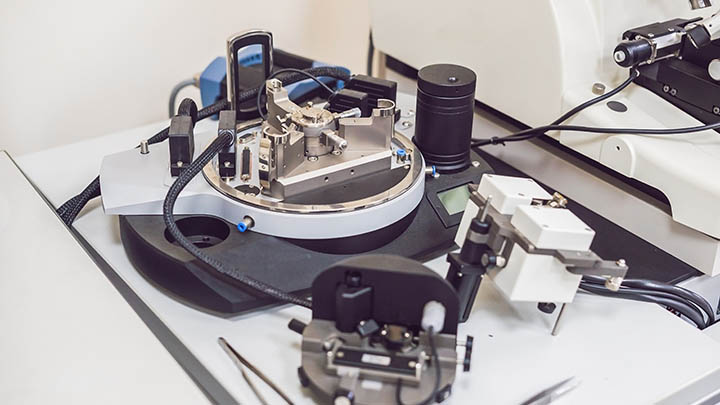

Technologies and equipment
What we offer
Cutting-edge imaging projects require access to the latest imaging equipment.
Constant investment in the Bioimaging Core Facility ensure that our equipment remains cutting-edge and allows our users access to the latest imaging techniques and technologies.
The facility is spread over three purpose-built suites in the Michael Smith and Stopford Buildings. We have five full-time members of staff who provide full training and support on a wide range of imaging techniques.
- Brucker Multimode 8
- Brucker Catalyst
- JPK Nanowizard
- Asylum Research MFP-3D
Atomic Force Microscopy (AFM) involves the raster scanning of a cantilever-bound probe over a surface and measures the deflection of the cantilever at each position to produce a 3D image of the sample with near atomic resolution.
In addition to imaging the AFM can also be used to perform force spectroscopy. Here the probe is moved in the Z direction towards the sample surface only, indenting slightly on contact before retracting again. The deflection of the cantilever is used to calculate the force applied to the probe at each displacement, along with adhesion forces as the cantilever retracts. In this way the forces involved in polymer folding/unfolding, biomolecular interactions and the compression of biomaterials (including cells) can be measured.
- Leica SP8 WLL inverted confocal
- Leica SP8 WLL inverted confocal with High Content Screening (HCS) capability
- Leica SP8 WLL upright confocal (dipping lenses, Clarity lens)
- Leica SP8 upright multi-photon capable of in vivo imaging
- Leica SP8 WLL 3x gSTED super resolution confocal with FLIM and FCS
- Zeiss LSM 880 inverted AiryScan confocal with FCS
- Zeiss LSM 880 inverted AiryScan confocal with multi-photon
- Zeiss LSM 880 upright AiryScan confocal
Confocal microscopy works by rastering a single diffraction limited laser spot across the sample so that excitation and detection is achieved “point by point” to build up the complete image. The use a confocal pinhole prevents out of focus light from reaching the detector and provides a complete 3D volume of the sample that is free from blur with a high level of resolution.
Super Resolution is achieved using Stimulated Emission Depletion (STED) microscopy. A scanning laser is used to excite the fluorophore as in a regular confocal system, but a second “doughnut” shaped laser is superimposed with this laser and causes a depletion of fluorescence inside that area. This STED effect takes the resolution below the diffraction limit for that wavelength and provides super-resolution images.
FCS/FLIM are techniques that allow the confocal system to measure protein interactions and dynamics by following changes in their mobility within the image beam (FCS) or the time the protein remains in the excited fluorescent state (FLIM).
Two-photon confocal microscopy requires two photons of long wavelength excitation light to arrive simultaneously at the fluorophore to cause excitation. The only place with a density of photons high enough for this to occur is at the focal plane and as a result there is no need for a confocal pinhole to reject out of focus light as there isn’t any out of focal plane excitation. This non-reliance on a pinhole to achieve optical sectioning and use of long wavelength excitation light makes the two-photon microscope ideally suited for deep tissue imaging and in vivo experiments.
- 3I Marianas Yokogawa CSU-X1 spinning disc confocal with galvo Vector FRAP head and FLIM
- 3I Marianas Yokogawa CSU-X1 spinning disc confocal
- Andor Dragonfly confocal with mosaic FRAP head
- Zeiss Z1 lightsheet
- Zeiss Z1 lightsheet for cleared samples (Clarity lens)
- 3I Lattice Light Sheet (LLS)
Spinning disk confocals use an array of micro-lenses to split the excitation laser light into many smaller beamlets that scan simultaneously across the sample. The emitted light then passes through a matched spinning disc of pinholes that reject the out of focus light and produce a confocal image. Since this technique uses hundreds of beamlets, rather than a single rastered laser, production of confocal images and z-stacks is therefore faster and less likely to cause photo-bleaching or cell damage than normal confocal, making spinning discs the ideal choice for confocal imaging of live cells.
Light Sheet microscopes uses an orthogonal arrangement of camera and illumination to create an 3D confocal stack of the samples. This is achieved using a thin sheet of laser light (1um to 14um thick) to illuminate just the focal plane of the objective, thus there is no need for a confocal pinhole as just fluorophores within the focal plane are illuminated. By moving the sample quickly through this optical arrangement of plane illumination and imaging, it is possible to achieve high speed confocal volumes with very low photobleaching.
- 2 x Nikon Long Term timelapse systems with JOBS modules
- PhaseFocus Livecyte
- Essen Incucyte S3
- Zeiss CellDiscoverer7 widefield with LSM 900 AiryScan 2 confocal
These systems are designed for maintaining cells for long periods of time by maintaining a stable environment of temperature, humidity, and CO2. Motorised XYZ stages, laser autofocus, and intelligent software allows multi-dimensional experiments to be carried out easily on multi-well plates and advanced analysis software is able to batch process these data sets to provide unbiased results on growth patterns, drug treatments and transfection studies. A range of imaging techniques are available from label free phase imaging, widefield fluorescence and AiryScan confocal imaging on whole plates over several days of imaging.
Snapshot
- 3 x Zeiss AxioImager Z1 fluorescence microscopes
- Leica M205 stereo fluorescence microscope
- Olympus BX63 with colour camera
Although these are the simplest microscope systems that we have, they are also our brightest and take great pictures so it may be that you don’t need to use the more complicated microscopes to answer your scientific questions. The microscopes are manual and very easy to use.
High end
- Olympus IX83 with MMI laser microdissection
- 2 x Olympus IX83 for deconvolution imaging
- Nikon Ti with Primo protein patterning
- Leica TIRF microscope with Infinity FRAP head
- Molecular Devices Flexstation3 fluorescence plate reader
Laser microdissection: uses a focused UV laser to cut around cells and small areas of tissue mounted on membrane slides. This cut tissue is then picked up using special sticky-cap Eppendorf tubes. Thus small regions of the samples can be cut and separated from the rest of the tissue without any physical handling of the sample and is suitable for downstream applications in the Genomic Technologies and Mass Spectroscopy core facilities.
Deconvolution: takes a series of widefield images through the sample, each of which contains both the in-focus light and out-of-focus information for each z-position. A mathematical algorithm is then performed on this data set to relocate the out of focus light back into its correct focal plane to generate sharp, bright, free from blur optical stack. However, since none of the light was rejected during the imaging process, the technique is more sensitive than regular confocal imaging and is ideal for samples where the fluorescence is weak.
Primo protein micro-patterning: allows the user to create any pattern and/or gradient of proteins on a coverslip based dish. These can then be seeded with cells and imaged on any of our microscope systems.
TIRF: Total Internal Reflection (TIRF) microscopyis a technique that allows the imaging of fluorophores very close to the coverslip while the rest of the sample remains “dark” producing high contrast images with very low photo-toxicity. The angle of the incident laser is adjusted to cause total internal reflection at the coverslip-sample buffer boundary. While this ensures that the sample is no longer excited by a penetrative laser beam, an evanescent illumination field is generated close to the coverslip on the sample side. This evanescent field excites any fluorophores within only a few 100nm of the coverslip while the remaining volume remains unexcited and thus permits imaging of proteins in contact (or very close proximity) to the coverslip.
Fluorescence plate reader: This system does not produce images but instead gives a fluorescent read out for each well. It can be used with 96 or 384 well plates to measure values for the following: Fluorescence Absorbance (200 to 1000nm), Fluorescence Emission Intensity (250nm to 850nm), Fluorescence Polarization, Time-Resolved Fluorescence and Luminescence. The system is temperature controlled and also has an 8 channel fluidics module for the automatic addition of drugs or reagents during the time-course.
Bioluminescence
- Olympus LV200 bioluminescence system
- Zeiss AxioObserver Z1 with motorised XY stage
Bioluminescence does not require an excitation light source in order for the protein of interest to glow. Instead, bioluminescent or chemiluminescent signals are produced within the cells/tissue. These can be imaged in live cells over very long time periods (hours-weeks). However, the weak signal emitted from these samples requires very long exposures and the system needs to be light tight to prevent ambient light from entering the system. The system has a 35mm dish holder/heater an environmental control chamber (for 37°C and 5% CO2) all contained in a light tight enclosure.
We have two 3D Histech Pannoramic P250 which can automatically detect and digitise up to 250 slides in a single run.
The slides can be brightfield or 4-channel fluorescence and images can be analysed using 3D Histech free slideviewer software or free software such as QuPath.
Our Olympus BX63 with scanning stage can be used to create high resolution images where the user is able to defined both the region of interest and its focus map.
This is then scanned and stitched into a single large image which can be opened in most free software such as QuPath and Fiji ImageJ.
Publications and outputs
Supporting high-impact publications
We are an essential resource that contributes to numerous high impact publications every year.
Here are some of the recent highlights.
Journal of Cell Science, 2021.
A high impact paper showcasing the range of microscopes available in the facility, from simple snapshot through to confocal microscopy and live cell imaging.
The paper also shows the power of combining light and EM microscopy.
- DOI: 10.1242/jcs.258549
Nature Communications, 2019.
Understanding how a complex system is co-ordinated and controlled requires a knowledge of the number of molecules involved.
This paper uses FCS to accurately predict this number of molecules and then image cells over several days to follow their fate.
- DOI: s41467-019-10734-8
Journal of Cell Science, 2020.
FCS, live cell imaging and modelling showcase the techniques available in the facility and how they can be combined to produce a high impact publication.
- DOI: 10.1242/jcs.248781
Journal of Cell Biology, 2019.
Combining live cell imaging and photokinetic capabilities of the systems to understand the mechanisms involved in cell adhesion and migration.
External access
Working with industry and other academic institutions
Although primarily focused on supporting research at The University of Manchester, we are also able to support work from other academic institutions and industry.
Please contact our Business Development Manager for further details if you are interested in accessing the facility
Dr Joanne Flannelly
Email: joanne.flannelly@manchester.ac.uk
Academic institutions
We are happy to provide access to all our microscope systems and slidescanning service to users from any academic institution.
We will provide these users with the same level of training and support as we would for our internal users.

Industry
We have worked for a number of multinational companies, both on an individual basis and as part of packages of work involving multiple facilities.
Work may be on a collaborative basis or purely as a service provision.
All work is fully documented and contractual, and non-disclosure compliance is assured.

Contact us
Find out more
Get in touch for further information or to inquire about using our facility.
Dr Peter March, Principal Technologist
Email: peter.march@manchester.ac.uk
Telephone: +44 (0)161 275 1571
Bioimaging Core Facility
The Michael Smith Building
The University of Manchester
Oxford Road
Manchester
M13 9PT
Maps and travel
We are based in the AV Hill Building (Building 71 on the University campus map).
Technology platforms
Technology platforms
We have a pioneering environment and facilities for research, innovation and technology development.
Technology platforms main page
